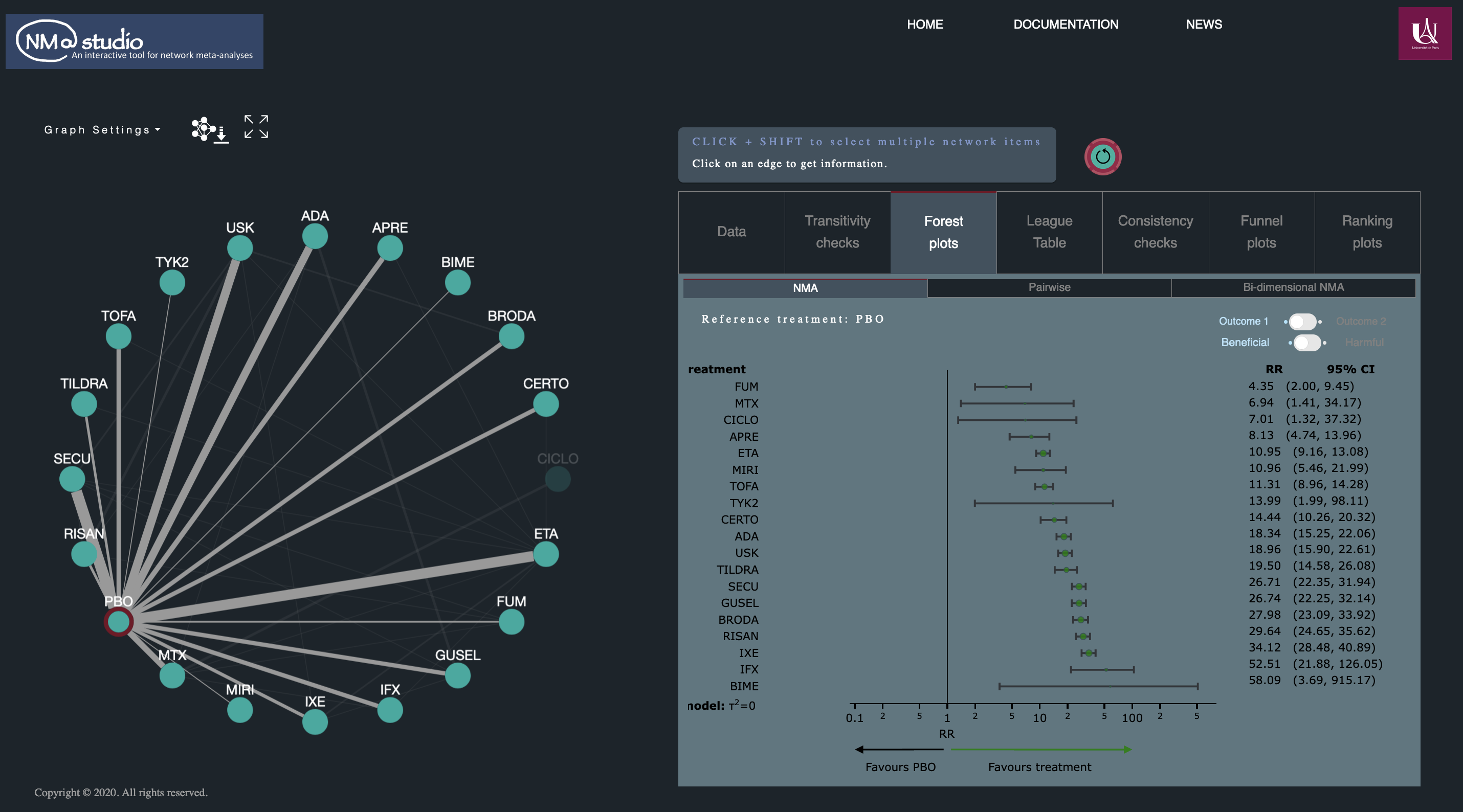
NMAstudio application
In network meta-analyses, visualisation of all the available evidence and interpretation of findings can be challenging, especially with networks of many treatments. NMAstudio is an open-source web application that facilitates the production and visualisation of NMA outputs in a fully interactive environment, providing easy ‘point-and-click’ interactions between the outputs and a network diagram. The software is fully built in Python but the R package netmeta is used to produce the analyses. Currently, the main features provided are: (a) boxplots of effect modifiers for the evaluation of transitivity; (b) pairwise or NMA forest plots and bi-dimensional plots for two outcomes; (c) league tables coloured by risk of bias or confidence ratings from CINeMA; (d) inconsistency tests and comparison-adjusted funnel plots for small-study effects; (e) ranking plots; (f) evolution of the network over time. All plots and tables are customisable and downloadable.
Citation:
Metelli S, Yu T, Chaimani A. NMAstudio: a fully interactive web-application for producing and visualising network meta-analyses. Annual Cochrane Colloquium 2023, London, UK.
Warning: Please, avoid using Firefox because some features might not work well. Preferred browsers are Safari and Chrome.
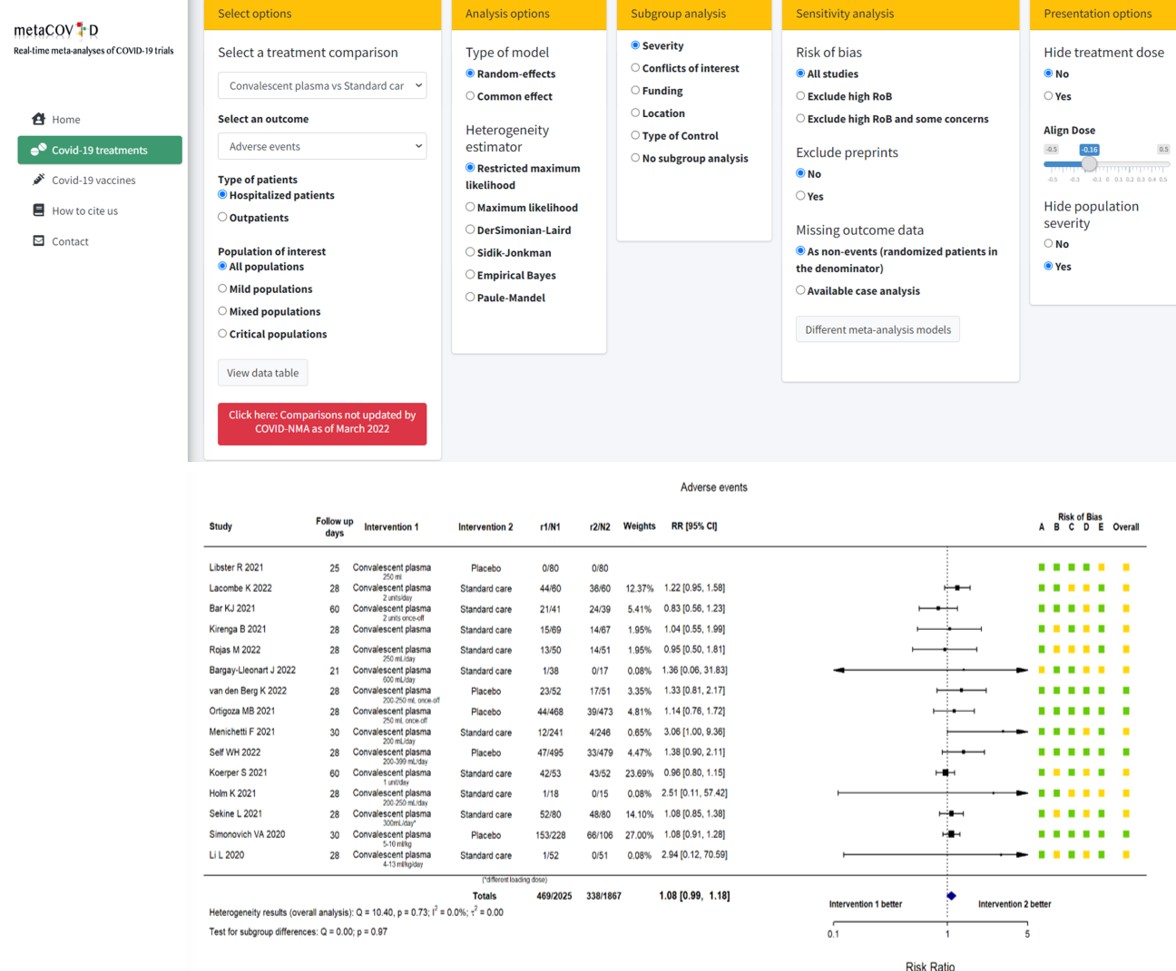
metaCOVID application
metaCOVID is a web-application that allows anyone interested in the results for COVID-19 treatments to directly use the most up-to-date database and perform their preferred meta-analyses in a user-friendly environment. It has been created in R and is freely available as an R-Shiny application. The application conducts living meta-analyses for 13 different outcomes and every available treatment comparison. Several options are available for subgroup and sensitivity analyses. The results are presented in downloadable forest plots. All produced forest plots include risk of bias assessments for every study as well as information on other study characteristics.
Citation:
Evrenoglou T, Boutron I, Seitidis G, Ghosn L, Chaimani A. metaCOVID: An R-Shiny application for living meta-analyses of COVID-19 trials. Res Synth Meth 2023;14:479–488
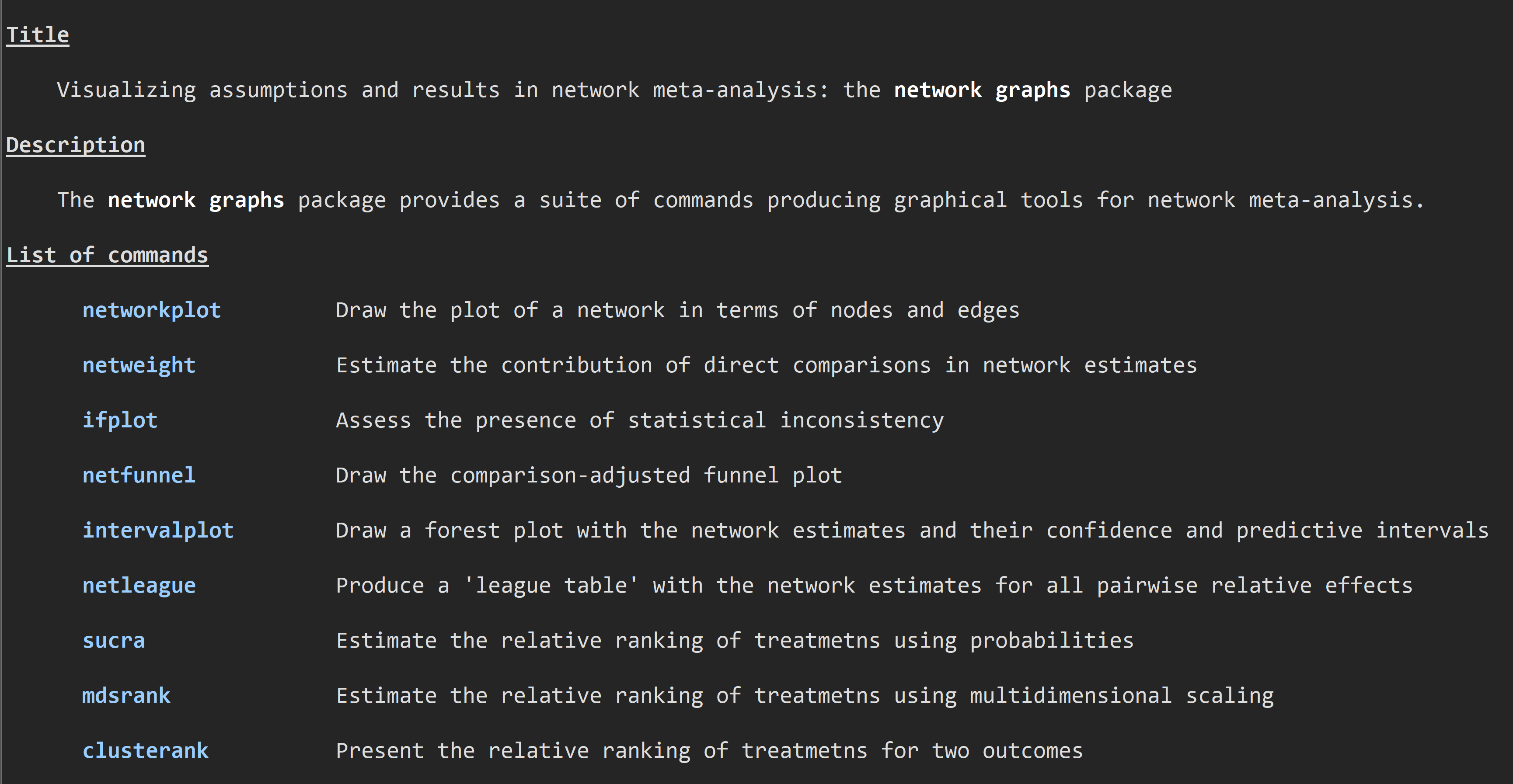
network graphs Stata package
Although network meta-analysis has been established as a useful evidence synthesis tool it has been often criticized for its complexity and for been accessible only to researchers with strong statistical and computational skills. Careful evaluation of its assumptions and understandable, concise presentation of the results are necessary to avoid misinterpretations and inform decision-making. We provide a series of Stata commands that can be used to produce useful graphical and numerical tools to enhance understanding of network meta-analysis procedures and findings. Latest update: 07 July 2019
Citations:
Chaimani A, Salanti G. Visualizing assumptions and results in network meta-analysis: the network graphs package. Stata Journal 2015; 15(4): 905-950.
Chaimani A, Higgins JP, Mavridis D, Spyridonos P, Salanti G. Graphical Tools for Network Meta-Analysis in STATA. PLoS One. 2013 Oct 3;8(10):e76654.
The package can be downloaded by typing in the command window:
net from https://www.cer-methods.com/Stata/
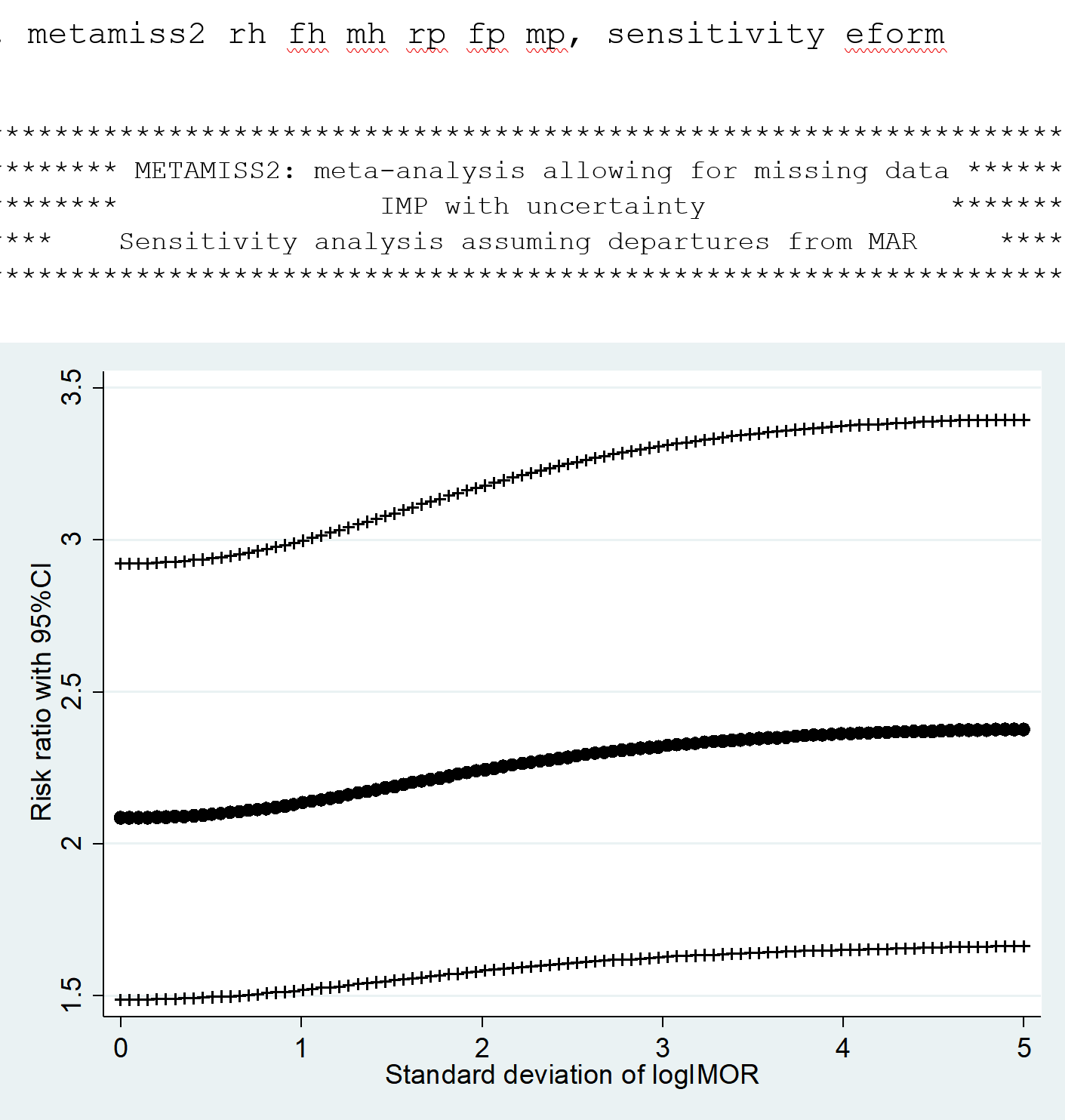
metamiss2 Stata package
Missing outcome data are a common threat to the validity of randomized trials and their meta-analysis, as they require making untestable assumptions. Researchers typically ignore missing data and analyze complete data only; this approach is equivalent to assuming that missing participants are missing at random. The metamiss2 command is based on the use of informative missingness parameters that relate the outcome in the missing data with that in the observed data. It performs a two-stage approach: it first estimates the ‘adjusted’ study-specific relative effects and their variances and covariances, and then calls metan or network meta to obtain the summary effects. Latest update: 01 October 2018.
Citation:
Chaimani A, Mavridis D, Salanti G, Higgins JPT, White IR. Allowing for informative missingness in aggregate data meta-analysis with continuous or binary outcomes: Extensions to metamiss. Stata J 2018; 18(3):716-740
The package can be downloaded by typing in the command window:
net from https://www.cer-methods.com/Stata/